library(tidyverse)
library(tidymodels)
library(pROC)
library(knitr)
library(kableExtra)
# set default theme in ggplot2
ggplot2::theme_set(ggplot2::theme_bw())Logistic Regression
Inference
Announcements
Statistics experience due TODAY at 11:59pm
HW 04 due Thursday, April 9 at 11:59pm
- released after class
Next project milestone: Draft report due Friday, April 10 (before lab)
Topics
Drop-in-deviance test
Inference for an individual coefficient
Computational setup
Risk of coronary heart disease
This data set is from an ongoing cardiovascular study on residents of the town of Framingham, Massachusetts. We want to examine the relationship between various health characteristics and the risk of having heart disease.
high_risk:- 1: High risk of having heart disease in next 10 years
- 0: Not high risk of having heart disease in next 10 years
age: Age at exam time (in years)totChol: Total cholesterol (in mg/dL)currentSmoker: 0 = nonsmoker, 1 = smokereducation: 1 = Some High School, 2 = High School or GED, 3 = Some College or Vocational School, 4 = College
Which model do we choose?
| term | estimate |
|---|---|
| (Intercept) | -6.673 |
| age | 0.082 |
| totChol | 0.002 |
| currentSmoker1 | 0.443 |
| term | estimate |
|---|---|
| (Intercept) | -6.456 |
| age | 0.080 |
| totChol | 0.002 |
| currentSmoker1 | 0.445 |
| education2 | -0.270 |
| education3 | -0.232 |
| education4 | -0.035 |
Log-Likelihood
Recall the log-likelihood function
\[ \begin{aligned} \log L&(\boldsymbol{\beta}|x_1, \ldots, x_n, y_1, \dots, y_n) \\ &= \sum\limits_{i=1}^n[y_i \log(\pi_i) + (1 - y_i)\log(1 - \pi_i)] \end{aligned} \]
where \(\pi_i = \frac{\exp\{\mathbf{x}_i^\mathsf{T}\boldsymbol{\beta}\}}{1 + \exp\{\mathbf{x}_i^\mathsf{T}\boldsymbol{\beta}\}}\)
Comparing the models using AIC and BIC
AIC
# AIC reduced model
glance(high_risk_fit_reduced)$AIC[1] 3232.812# AIC full model
glance(high_risk_fit_full)$AIC[1] 3231.6BIC
# BIC reduced model
glance(high_risk_fit_reduced)$BIC[1] 3258.074# BIC full model
glance(high_risk_fit_full)$BIC[1] 3275.807Drop-in-deviance test
Drop-in-deviance test
We can use the drop-in-deviance test (aka Likelihood Ratio Test) to test
the overall statistical significance of a logistic regression model
the statistical significance of a subset of coefficients in the model
Deviance
The deviance is a measure of the degree to which the predicted values are different from the observed values (compares the current model to a “saturated” model)
In logistic regression,
\[ D = -2 \log L \]
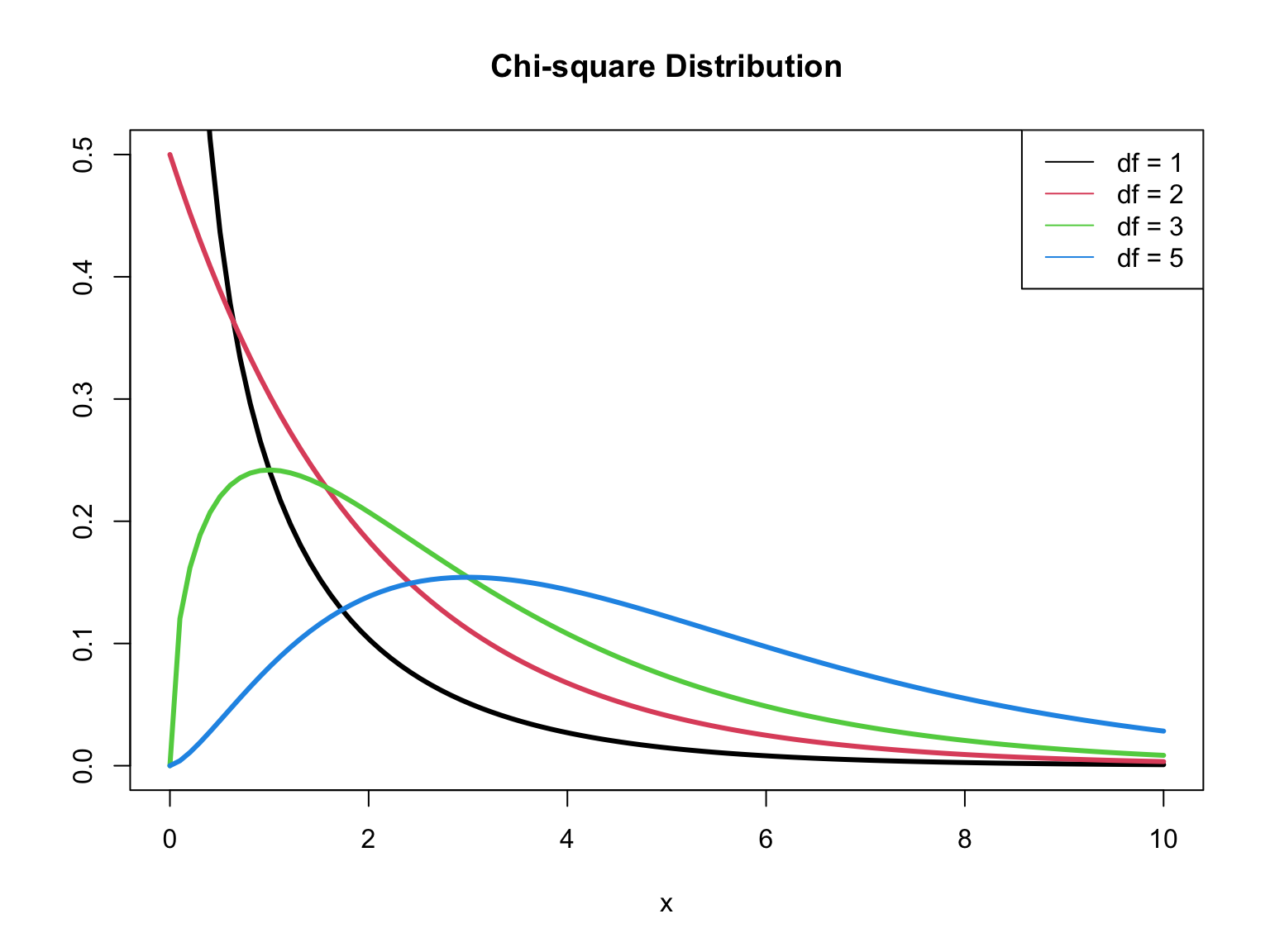
\(D \sim \chi^2_{n - p - 1}\) (\(D\) follows a Chi-square distribution with \(n - p - 1\) degrees of freedom)1
Note: \(n - p - 1\) = the degrees of freedom associated with the error in the model (like residuals)
\(\chi^2\) distribution
Test for overall significance
We can test the overall significance for a logistic regression model, i.e., whether there is at least one predictor with a non-zero coefficient
\[ \begin{aligned} &H_0: \beta_1 = \dots = \beta_p = 0 \\ &H_a: \beta_j \neq 0 \text{ for at least one } j \end{aligned} \]
. . .
The drop-in-deviance test for overall significance compares the fit of a model with no predictors to the current model.
Drop-in-deviance test statistic
Let \(L_0\) and \(L_a\) be the likelihood functions of the model under \(H_0\) and \(H_a\), respectively. The test statistic is
\[ \begin{aligned} G = D_0 - D_a &= (-2\log L_0) - (-2\log L_a)\\[5pt] & = -2(\log L_0 - \log L_a) \\[5pt] &= -2\sum_{i=1}^n \Big[ y_i \log \Big(\frac{\hat{\pi}^0}{\hat{\pi}^a_i}\Big) + (1 - y_i)\log \Big(\frac{1-\hat{\pi}^0}{1-\hat{\pi}^a_i}\Big)\Big] \end{aligned} \]
where \(\hat{\pi}^0\) is the predicted probability under \(H_0\) and \(\hat{\pi}_i^a = \frac{\exp \{\mathbf{x}_i^\mathsf{T}\boldsymbol{\beta}\}}{1 + \exp \{\mathbf{x}_i^\mathsf{T}\boldsymbol{\beta}\}}\) is the predicted probability under \(H_a\) 2
Drop-in-deviance test statistic
\[ G = -2\sum_{i=1}^n \Big[ y_i \log \Big(\frac{\hat{\pi}^0}{\hat{\pi}^a_i}\Big) + (1 - y_i)\log \Big(\frac{1-\hat{\pi}^0}{1-\hat{\pi}^a_i}\Big)\Big] \]
. . .
When \(n\) is large, \(G \sim \chi^2_p\), ( \(G\) follows a Chi-square distribution with \(p\) degrees of freedom)
The p-value is calculated as \(P(\chi^2 > G)\)
Large values of \(G\) (small p-values) indicate at least one \(\beta_j\) is non-zero
Heart disease model: drop-in-deviance test
\[ \begin{aligned} &H_0: \beta_{age} = \beta_{totChol} = \beta_{currentSmoker} = 0 \\ &H_a: \beta_j \neq 0 \text{ for at least one }j \end{aligned}\]
. . .
Fit the null model (we’ve already fit the alternative model)
null_model <- glm(high_risk ~ 1, data = heart_disease, family = "binomial")| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -1.72294 | 0.0436342 | -39.486 | 0 |
Heart disease model: drop-in-deviance test
Compute the log-likelihoods
(L_0 <- glance(null_model)$logLik)[1] -1737.735(L_a <- glance(high_risk_fit)$logLik)[1] -1612.406. . .
Compute the test statistic
(G <- -2 * (L_0 - L_a))[1] 250.6572. . .
Heart disease model: drop-in-deviance test
Compute the p-value
pchisq(G, df = 3, lower.tail = FALSE)[1] 4.717158e-54. . .
Conclusion
The p-value is small, so we reject \(H_0\). The data provide evidence that at least one predictor in the model has a non-zero coefficient.
Why use overall test?
Why do we use a test for overall significance instead of just looking at the test for individual coefficients?3
. . .
Suppose we have a model such that \(p = 100\) and \(H_0: \beta_1 = \dots = \beta_{100} = 0\) is true
. . .
About 5% of the p-values for individual coefficients will be below 0.05 by chance.
So we expect to see 5 small p-values if even no linear association actually exists.
Therefore, it is very likely we will see at least one small p-value by chance.
The overall test of significance does not have this problem. There is only a 5% chance we will get a p-value below 0.05, if a relationship truly does not exist.
Test a subset of coefficients
Testing a subset of coefficients
Suppose there are two models:
Reduced Model: includes predictors \(x_1, \ldots, x_q\)
Full Model: includes predictors \(x_1, \ldots, x_q, x_{q+1}, \ldots, x_p\)
We can use a drop-in-deviance test to determine if any of the new predictors are useful
. . .
\[ \begin{aligned} &H_0: \beta_{q+1} = \dots = \beta_p = 0\\ &H_a: \beta_j \neq 0 \text{ for at least one }j \end{aligned} \]
Drop-in-deviance test
\[ \begin{aligned} &H_0: \beta_{q+1} = \dots = \beta_p = 0\\ &H_a: \beta_j \neq 0 \text{ for at least one }j \end{aligned} \]
. . .
The test statistic is
\[ \begin{aligned} G = D_{reduced} - D_{full} &= (-2\log L_{reduced}) - (-2 \log L_{full}) \\ &= -2(\log L_{reduced} - \log L_{full}) \end{aligned} \]
. . .
The p-value is calculated using a \(\chi_{\Delta df}^2\) distribution, where \(\Delta df\) is the number of parameters being tested (the difference in number of parameters between the full and reduced model).
Example: Include education?
Should we include education in the model?
Reduced model:
age,totChol,currentSmokerFull model:
age,totChol,currentSmoker,education
. . .
\[ \begin{aligned} &H_0: \beta_{ed2} = \beta_{ed3} = \beta_{ed4} = 0 \\ &H_a: \beta_j \neq 0 \text{ for at least one }j \end{aligned} \]
Example: Include education?
reduced_model <- glm(high_risk ~ age + totChol + currentSmoker,
data = heart_disease, family = "binomial")
full_model <- glm(high_risk ~ age + totChol + currentSmoker + education,
data = heart_disease, family = "binomial"). . .
Compute deviances
(deviance_reduced <- -2 * glance(reduced_model)$logLik)[1] 3224.812(deviance_full <- -2 * glance(full_model)$logLik)[1] 3217.6. . .
Compute test statistic
(G <- deviance_reduced - deviance_full)[1] 7.212113Example: Include education?
Compute p-value
pchisq(G, df = 3, lower.tail = FALSE)[1] 0.06543567. . .
What is your conclusion? Would you include education in the model that already has age, totChol, currentSmoker?
Let’s take a look at education
| term | estimate | std.error | statistic | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|
| (Intercept) | -6.456 | 0.391 | -16.533 | 0.000 | -7.230 | -5.698 |
| age | 0.080 | 0.006 | 13.571 | 0.000 | 0.068 | 0.091 |
| totChol | 0.002 | 0.001 | 2.086 | 0.037 | 0.000 | 0.004 |
| currentSmoker1 | 0.445 | 0.094 | 4.747 | 0.000 | 0.262 | 0.630 |
| education2 | -0.270 | 0.113 | -2.379 | 0.017 | -0.494 | -0.049 |
| education3 | -0.232 | 0.135 | -1.714 | 0.086 | -0.501 | 0.029 |
| education4 | -0.035 | 0.150 | -0.232 | 0.817 | -0.335 | 0.255 |
What is your final assessment about including education in the model?
Drop-in-deviance test in R
Conduct the drop-in-deviance test using the anova() function in R with option test = "Chisq"
anova(reduced_model, full_model, test = "Chisq") |>
tidy() |>
kable(digits = 3)| term | df.residual | residual.deviance | df | deviance | p.value |
|---|---|---|---|---|---|
| high_risk ~ age + totChol + currentSmoker | 4082 | 3224.812 | NA | NA | NA |
| high_risk ~ age + totChol + currentSmoker + education | 4079 | 3217.600 | 3 | 7.212 | 0.065 |
Add interactions with currentSmoker?
| term | df.residual | residual.deviance | df | deviance | p.value |
|---|---|---|---|---|---|
| high_risk ~ age + totChol + currentSmoker | 4082 | 3224.812 | NA | NA | NA |
| high_risk ~ age + totChol + currentSmoker + currentSmoker * age + currentSmoker * totChol | 4080 | 3222.377 | 2 | 2.435 | 0.296 |
Inference for an individual coefficient
Distribution of \(\hat{\boldsymbol{\beta}}\)
When \(n\) is large, \(\hat{\boldsymbol{\beta}}\) is normally distributed
How do we know \(\hat{\boldsymbol{\beta}}\) is normally for large \(n\)?
Properties of \(\hat{\boldsymbol{\beta}}\)
\(\hat{\boldsymbol{\beta}}\) is a consistent estimator
\[ \lim_{n \rightarrow \infty} P(|\hat{\boldsymbol{\beta}} - \boldsymbol{\beta}| \geq c) = 0 \hspace{8mm} \forall ~ c > 0 \]
. . .
When \(n\) is large, \(Var(\hat{\boldsymbol{\beta}})\), the matrix of variances and covariances between estimators
\[ Var(\hat{\boldsymbol{\beta}}) = (\mathbf{X}^\mathsf{T}\mathbf{V}\mathbf{X})^{-1} \]
where \(\mathbf{V}\) is a \(n\times n\) diagonal matrix, such that \(V_{ii}\) is the estimated variance for the \(i^{th}\) observation
Test for a single coefficient
Hypotheses: \(H_0: \beta_j = 0 \hspace{2mm} \text{ vs } \hspace{2mm} H_a: \beta_j \neq 0\), given the other variables in the model
. . .
(Wald) Test Statistic: \[z = \frac{\hat{\beta}_j - 0}{SE(\hat{\beta}_j)}\]
where \(SE(\hat{\beta}_j)\) is the square root of the \(j^{th}\) diagonal element of \(Var(\hat{\boldsymbol{\beta}})\)
. . .
P-value: \(P(|Z| > |z|)\), where \(Z \sim N(0, 1)\), the Standard Normal distribution
Coefficient for age
| term | estimate | std.error | statistic | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|
| (Intercept) | -6.673 | 0.378 | -17.647 | 0.000 | -7.423 | -5.940 |
| age | 0.082 | 0.006 | 14.344 | 0.000 | 0.071 | 0.094 |
| totChol | 0.002 | 0.001 | 1.940 | 0.052 | 0.000 | 0.004 |
| currentSmoker1 | 0.443 | 0.094 | 4.733 | 0.000 | 0.260 | 0.627 |
. . .
Hypotheses:
\[ H_0: \beta_{age} = 0 \hspace{2mm} \text{ vs } \hspace{2mm} H_a: \beta_{age} \neq 0 \], given total cholesterol and smoking status are in the model.
Coefficient for age
| term | estimate | std.error | statistic | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|
| (Intercept) | -6.673 | 0.378 | -17.647 | 0.000 | -7.423 | -5.940 |
| age | 0.082 | 0.006 | 14.344 | 0.000 | 0.071 | 0.094 |
| totChol | 0.002 | 0.001 | 1.940 | 0.052 | 0.000 | 0.004 |
| currentSmoker1 | 0.443 | 0.094 | 4.733 | 0.000 | 0.260 | 0.627 |
Test statistic:
\[z = \frac{ 0.0825 - 0}{0.00575} = 14.34 \]
Coefficient for age
| term | estimate | std.error | statistic | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|
| (Intercept) | -6.673 | 0.378 | -17.647 | 0.000 | -7.423 | -5.940 |
| age | 0.082 | 0.006 | 14.344 | 0.000 | 0.071 | 0.094 |
| totChol | 0.002 | 0.001 | 1.940 | 0.052 | 0.000 | 0.004 |
| currentSmoker1 | 0.443 | 0.094 | 4.733 | 0.000 | 0.260 | 0.627 |
P-value:
\[P(|Z| > |14.34|) \approx 0 \]
. . .
2 * pnorm(14.34,lower.tail = FALSE)[1] 1.230554e-46Coefficient for age
| term | estimate | std.error | statistic | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|
| (Intercept) | -6.673 | 0.378 | -17.647 | 0.000 | -7.423 | -5.940 |
| age | 0.082 | 0.006 | 14.344 | 0.000 | 0.071 | 0.094 |
| totChol | 0.002 | 0.001 | 1.940 | 0.052 | 0.000 | 0.004 |
| currentSmoker1 | 0.443 | 0.094 | 4.733 | 0.000 | 0.260 | 0.627 |
Conclusion:
The p-value is very small, so we reject \(H_0\). The data provide sufficient evidence that age is a statistically significant predictor of whether someone is high risk of having heart disease, after accounting for total cholesterol and smoking status.
Confidence interval for \(\beta_j\)
We can calculate the C% confidence interval for \(\beta_j\) as the following:
\[ \Large{\hat{\beta}_j \pm z^* \times SE(\hat{\beta}_j)} \]
where \(z^*\) is calculated from the \(N(0,1)\) distribution
. . .
This is an interval for the change in the log-odds for every one unit increase in \(x_j\)
Interpretation in terms of the odds
The change in odds for every one unit increase in \(x_j\).
\[ \Large{\exp\{\hat{\beta}_j \pm z^* \times SE(\hat{\beta}_j)\}} \]
. . .
Interpretation: We are \(C\%\) confident that for every one unit increase in \(x_j\), the odds multiply by a factor of \(\exp\{\hat{\beta}_j - z^* \times SE(\hat{\beta}_j)\}\) to \(\exp\{\hat{\beta}_j + z^* \times SE(\hat{\beta}_j)\}\), holding all else constant.
CI for age
| term | estimate | std.error | statistic | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|
| (Intercept) | -6.673 | 0.378 | -17.647 | 0.000 | -7.423 | -5.940 |
| age | 0.082 | 0.006 | 14.344 | 0.000 | 0.071 | 0.094 |
| totChol | 0.002 | 0.001 | 1.940 | 0.052 | 0.000 | 0.004 |
| currentSmoker1 | 0.443 | 0.094 | 4.733 | 0.000 | 0.260 | 0.627 |
Interpret the 95% confidence interval for age in terms of the odds of being high risk for heart disease.
Overview of testing coefficients
Test a single coefficient
Drop-in-deviance test
Wald hypothesis test and confidence interval
. . .
Test a subset of coefficients
- Drop-in-deviance test
. . .
Can use AIC and BIC to compare models in both scenarios
Recap
Model selection for logistic regression using a drop-in-deviance test
Inference for an individual coefficient
Next class
Data science ethics
No prepare assignment